August 8, 2011 - By Susan Young

Manuel Amieva
If you were the size of a bacterium, the lining of a stomach would seem like a rugged, hilly landscape filled with acid-spewing geysers, said Manuel Amieva, MD, PhD, assistant professor of pediatrics and of microbiology and immunology. Stomach-dwelling bacteria called Helicobacter pylori, the cause of ulcers and some gastric cancers, must navigate through the treacherous terrain to find sanctuary in the mucous layer that coats the inside of the stomach.
But now Amieva and his colleagues have identified a protein that regulates H. pylori’s ability to seek shelter. The protein could be a target for therapies that specifically combat H. pylori while leaving our friendly gut bacteria alone. The findings were published on July 26 in the journal mBio.
H. pylori makes a living in about half the people on Earth. For most, the tiny tenants go unnoticed. But in about 15 percent of those infected, the microbes cause ulcers. Yet worse, for others, it increases the risk of gastric cancer.
Despite the fact that it lives in our stomachs, H. pylori is a not fan of our caustic digestive acids. To avoid getting burned, the bacteria use their propeller-like appendages called flagella to power their corkscrew bodies through the mucous that protects our stomach cells. While it was known that the bacteria colonize the stomach’s surface mucus, the researchers found that they also take up residence deep in some of the glands that tunnel down from the stomach’s surface.
The team made the discovery while studying a protein called ChePep. Michael Howitt, PhD, a former graduate student in Amieva’s lab, created a special strain of H. pylori that cannot make the protein. Bacteria lacking ChePep don’t make their way down into the glands, prompting Howitt to examine how the bugs move.
Although the mutant bacteria could still swim, they behaved erratically. “They weren’t taking the nice arcing trajectories that you see with normal H. pylori, they were moving herky jerky, taking short swims, then stopping, then turning and swimming off again,” said Howitt, who is lead author of the study. The bugs were doing a lot more backward swimming than typical H. pylori, as if they were always trying to escape toxic conditions.
Even simple organisms like bacteria can sense and respond to their environment. Microbes use their so-called chemotaxis machinery to move based on the chemical composition of their surroundings, swimming toward good conditions and away from bad. “Bacteria don’t swim aimlessly and their ability to sense the environment is critical to survival,” said Howitt. The zone of safety for H. pylori is constantly changing as the mucous layer gets sloughed off.
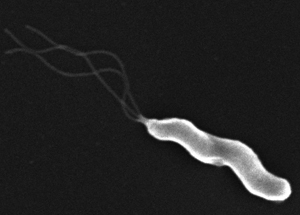
Scientists have found a way to disrupt Helicobacter pylori’s ability to use its whip-like flagella to move around the stomach. The bacteria causes ulcers and gastric cancer.
The microbes can change their direction by changing the spin of their flagella, the whip-like structures on one end of the microbes. “If the bacterium isn’t sensing anything bad, it’ll rotate the flagella in one direction and swim straight,” said Amieva, senior author of the study. “But if something bad is seen, it’ll activate a clutch, stall the flagella and switch directions.” Through genetic and microscopic analysis, the scientists concluded that ChePep helps to regulate the bacteria’s “clutch,” and thus is involved in changing the direction that the flagellum twirls.
“When other chemotaxis proteins are mutated, H. pylori just swims straight, like hands off the wheel, but ChePep mutants do the opposite, they look like they are constantly slamming the brake and turning,” said Howitt.
The researchers were surprised to find an unknown part to the chemotaxis machinery. “The chemotaxis system is the best-studied biochemical signaling system ever, so finding a new player was totally unexpected,” said Amieva. Much of what scientists know about the chemotaxis machinery of bacteria is based on just two bacteria species that aren’t that closely related to H. pylori. Nonetheless, much of the chemotaxis machinery is shared amongst most bacterial families. ChePep, however, is unique.
H. pylori belongs to a class of bacteria that thrive in extreme environments, like deep-sea hydrothermal vents, sulfurous caves and the guts of birds and mammals. “The epsilon proteobacteria do a good job at making a living in some inhospitable places,” said Howitt. The researchers found ChePep proteins in other epsilon proteobacteria and the other ChePep’s were so similar to H. pylori’s that Howitt could switch a copy from a deep-sea species into the stomach bugs with no observable difference.
However, the team did not find ChePep outside the epsilon proteobacteria, suggesting that disrupting the protein could be a way to specifically target H. pylori. “When you block chemotaxis, H. pylori is more sensitive to antibiotics,” said Howitt. “If you blocked ChePep, the beneficial bacteria in our guts would still have their chemotaxis machinery intact.”
In addition to Amieva and Howitt, graduate student Josephine Lee, microscopist Lydia-Marie Joubert and colleagues at UC-Santa Cruz and the Universitätsmedizin Mannheim in Germany contributed to the study. The work was funded by the NIH Ruth L. Kirschtein National Research Service Award, the National Cancer Institute and the National Institute of Allergy and Infectious Diseases. The departments of Pediatrics and of Microbiology and Immunology also supported this research.
Susan L. Young is a science-writing intern in the Office of Communication & Public Affairs.
About Stanford Medicine
Stanford Medicine is an integrated academic health system comprising the Stanford School of Medicine and adult and pediatric health care delivery systems. Together, they harness the full potential of biomedicine through collaborative research, education and clinical care for patients. For more information, please visit med.stanford.edu.